MCAW-DB |
Ver 3.0 updated July 10, 2019 |
MCAW-DB is a database whereby users can view the multiple alignment analysis results obtained from the MCAW tool [1], which can computationally find common glycan substructures among a group of glycan structures. The data in MCAW-DB are analysis results of glycan array experiments; those glycans having higher binding affinity to glycan binding proteins (GBPs) have been analyzed and can be viewed as “glycan recognition profiles”. Currently, this database provides published analysis results of glycan array data from the Consortium for Functional Glycomics (CFG).
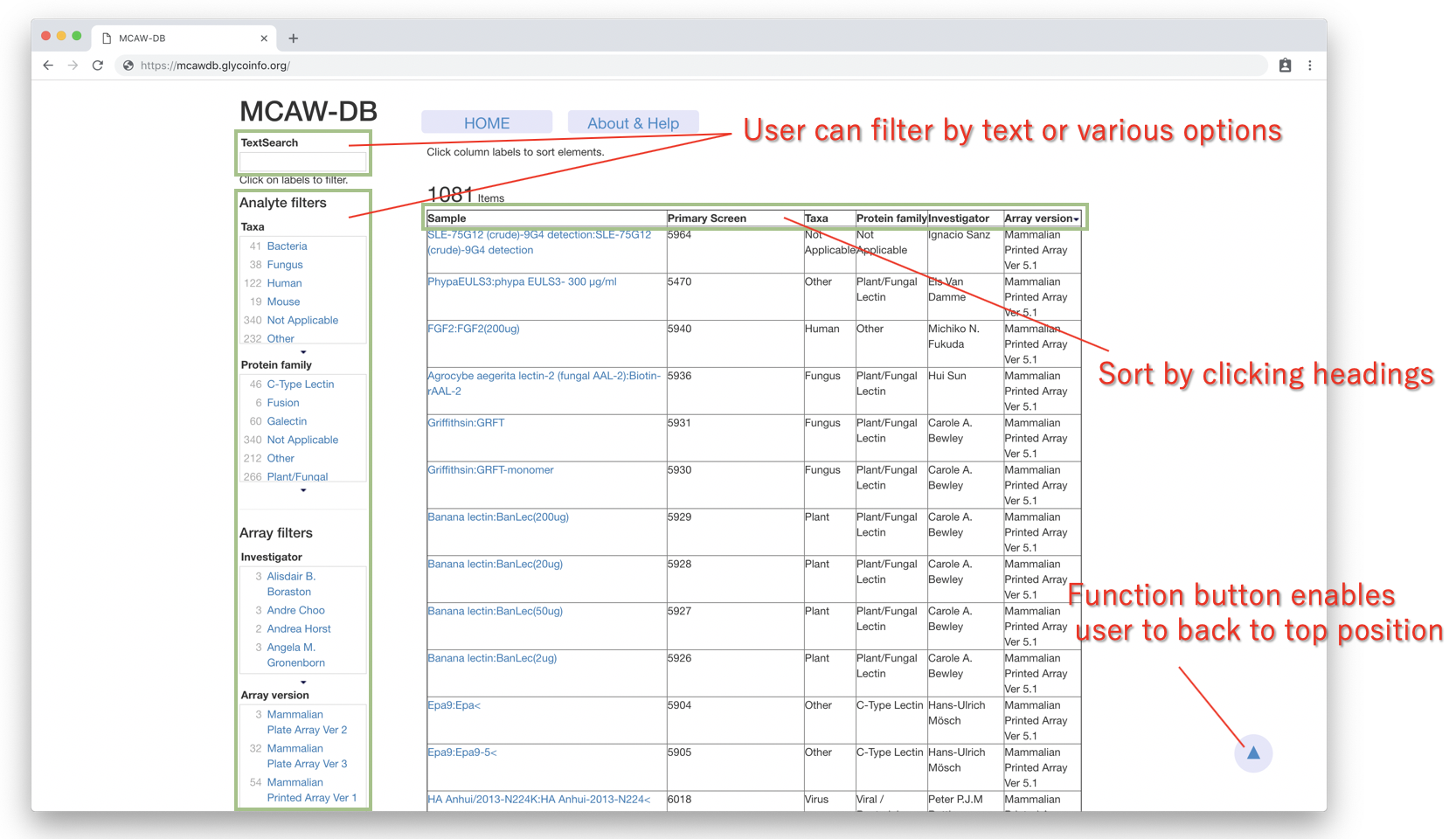
On the “MCAW-DB home screen”, users can filter by various options such as Species/Scientific Name protein family, investigator and array version.
In addition, text box enbles users to search by text, and the coverage is the same as listed above.
Users can view the each analysis result by clicking the sample name. A new “Analysis details page” will be displayed,
where the multiple alignment result, details about the experimental sample, and link to the original CFG data are provided.

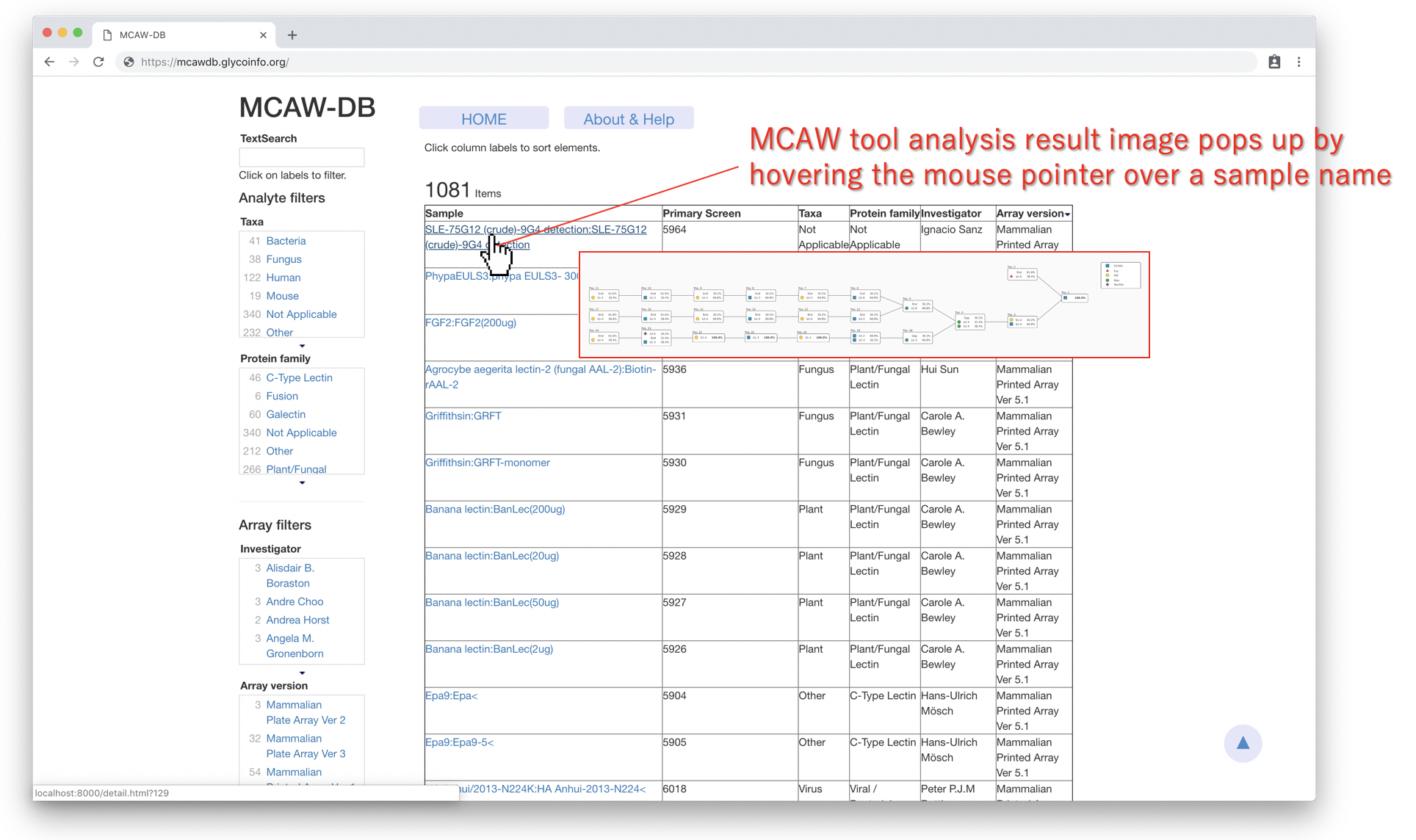
MCAW tool analysis result image would appear when the user hover the mouse cursor over the sample name of a sample. The Sturucture image would be possible to see bigger in the detail page.
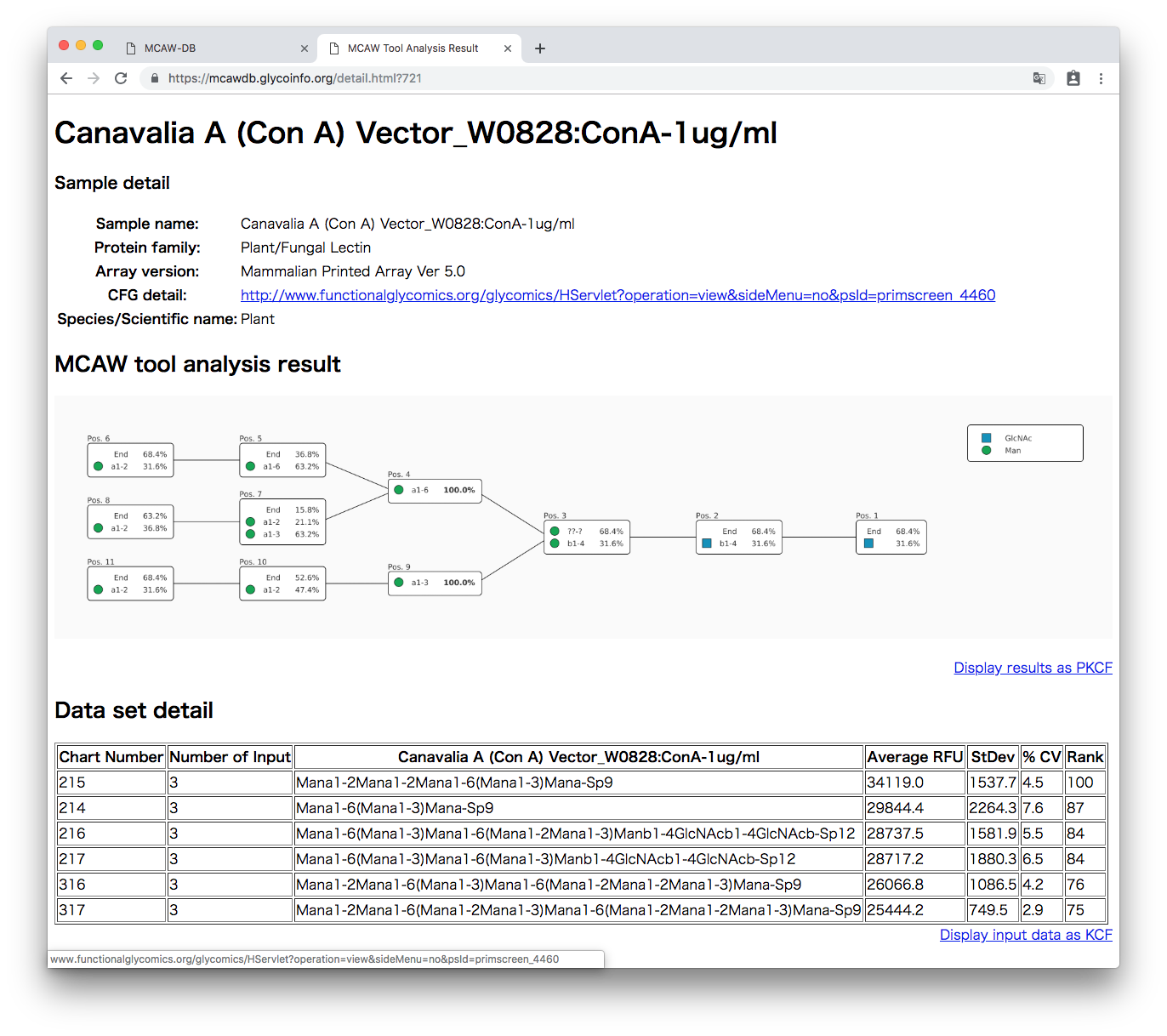
The alignment results are displayed using SNFG symbols for the monosaccharides.
The percentage of each node aligned at each position is listed. For example, in this Figure, position 1 (Pos.1) is 68.4% empty due to the End. Otherwise, GlcNAc appears 31.6%.
The link “Display result as PKCF” will display the alignment results in the text format called PKCF [2].
The bottom of the alignment results is a table of the input data. Each glycan is listed with their weights used when the analysis was executed.
MCAW-DB has been tested on the following: Internet Explorer 11, Firefox Quantum version 59, Google Chrome version 64 on Microsoft Windows 7, and Safari version 11, Firefox version 59 and Google Chrome version 64 on Mac OS. If MCAW-DB doesn't seem to display correctly on these environments, the user may need to reload the page due to data loading delays.
[1] Hosoda, M., Akune, Y., & Aoki-Kinoshita, K. F. (2017). Development and application of an algorithm to compute weighted multiple glycan alignments. Bioinformatics (Oxford, England), 33(9), 1317.
[2] Hosoda, M., Akune, Y., & Aoki-Kinoshita, K. F. (2012). Multiple tree alignment with weights applied to carbohydrates to extract binding recognition patterns. In IAPR International Conference on Pattern Recognition in Bioinformatics (pp. 49-58). Springer, Berlin, Heidelberg.
[3] Hosoda, M., Takahashi, Y., Shiota, M., Shinmachi, D., Inomoto, R., Higashimoto, S., & Aoki-Kinoshita, K. F. (2018). MCAW-DB: A glycan profile database capturing the ambiguity of glycan recognition patterns. Carbohydrate research, 464, 44-56.